

| Key Learning Goals for this Lesson: |
Textbook reading: Consult Course Schedule |
A quick review of regression, expectation, variance, and parameter estimation.
Input vector: \( X = (X_1, X_2, ... , X_p)\).
Output Y is real-valued.
Predict Y from X by f(X) so that the expected loss function \(E(L(Y, f(X)))\) is minimized.
Intuitively, the expectation of a random variable is its "average" value under its distribution.
Formally, the expectation of a random variable X, denoted E[X], is its Lebesgue integral with respect to its distribution.
If X takes values in some countable numeric set \(\chi\), then
\[E(X) =\sum_{x \in \chi}xP(X=x) \]
If \(X \in \mathbb{R}^m\) has a density p, then
\[E(X) =\int_{\mathbb{R}^m}xp(x)dx \]
Expectation is linear: \(E(aX +b)=aE(X) + b\).
Also, \(E(X+Y) = E(X) +E(Y)\).
Expectation is monotone: if X ≥Y, then E(X) ≥ E(Y).
The variance of a random variable X is defined as:
\(Var(X) = E[(X-E[X])^2]=E[X^2]-(E[X])^2\)
and the variance obeys the following \(a, b \in \mathbb{R}\):
\(Var(aX + b) =a^2Var(X)\)
The data x1, ... , xn is generally assumed to be independent and identically distributed (i.i.d.).
We would like to estimate some unknown value θ associated with the distribution from which the data was generated.
In general, our estimate will be a function of the data (i.e., a statistic)
\[\hat{\theta} =f(x_1, x_2, ... , x_n)\]
Example: Given the results of n independent flips of a coin, determine the probability p with which it lands on heads.
In practice, we often seek to select a distribution (model) corresponding to our data.
If our model is parameterized by some set of values, then this problem is that of parameter estimation.
How can we obtain estimates in general? One Answer: Maximize the likelihood and the estimate is called the maximum likelihood estimate, MLE.
\( \begin {align} \hat{\theta} & = argmax_{\theta} \prod_{i=1}^{n}p_{\theta}(x_i) \\
& =argmax_{\theta} \sum_{i=1}^{n}log (p_{\theta}(x_i)) \\
\end {align} \)
Discussion
Let's look at the setup for linear regression. We have an input vector: X = (X1, X2, ..., Xp ). This vector is p dimensional.
The output Y is a real value and is ordered.
We want to predict Y from X.
Before we actually do the prediction we have to train the function f(X). By the end of the training, I would have a function f(X) to map every X into an estimated Y. Then, we need some way to measure how good this predictor function is. This is measured by the expectation of a loss.
![]() Why do we have loss in the estimation?
Why do we have loss in the estimation?
Y is actually a random variable given X. For instance, consider predicting someones weight based on the person's height. People can have different weights given the same height. If you think of the weight as Y and the height as X, Y is random given X. We therefore cannot have a perfect prediction for every subject because f(X) is a fixed function, impossible to be correct all the time. The loss measures how different the true Y is from your prediction.
![]() Why do we have the overall loss expressed as an expectation?
Why do we have the overall loss expressed as an expectation?
The loss may be different for different subjects. In statistics, a common thing to do is to average the losses over the entire population.
Squared loss:
L(Y, f(X)) = (Y - f(X))2 .
We simply measure the difference between the two variables and square them so that we can handle negative and positive difference symmetrically.
Suppose the distribution of Y given X is known, the optimal predictor is:
f*(X) = argminf(X) E(Y - f(X))2
= E(Y | X) .
This is the conditional expectation of Y given X. The function E(Y | X) is called the regression function.
We want to predict the number of physicians in a metropolitan area.
Problem: The number of active physicians in a Standard Metropolitan Statistical Area (SMSA), denoted by Y, is expected to be related to total population (X1, measured in thousands), land area (X2, measured in square miles), and total personal income (X3, measured in millions of dollars). Data are collected for 141 SMSAs, as shown in the following table.
|
i :
|
1
|
2
|
3
|
...
|
139
|
140
|
141
|
|
X1
|
9387
|
7031
|
7017
|
...
|
233
|
232
|
231
|
|
X2
|
1348
|
4069
|
3719
|
...
|
1011
|
813
|
654
|
|
X3
|
72100
|
52737
|
54542
|
...
|
1337
|
1589
|
1148
|
|
Y
|
25627
|
15389
|
13326
|
...
|
264
|
371
|
140
|
Our Goal: To predict Y from X1, X2, and X3.
This is a typical regression problem.
\[ f(X)=\beta_{0} + \sum_{j=1}^{p}X_{j}\beta_{j}\]
This is just a linear combination of the measurements that are used to make predictions, plus a constant, (the intercept term). This is a simple approach. However, It might be the case that the regression function might be pretty close to a linear function, and hence the model is a good approximation.
![]() What if the model is not true?
What if the model is not true?
Comments on Xj :
Below is a geometric interpretation of a linear regression.
For instance, if we have two variables, X1 and X2, and we predict Y by a linear combination of X1 and X2, the predictor function corresponds to a plane (hyperplane) in the three-dimensional space of X1, X2, Y. Given a pair of X1 and X2 we could find the corresponding point on the plane to decide Y by drawing a perpendicular line to the hyperplane, starting from the point in the plane spanned by the two predictor variables.
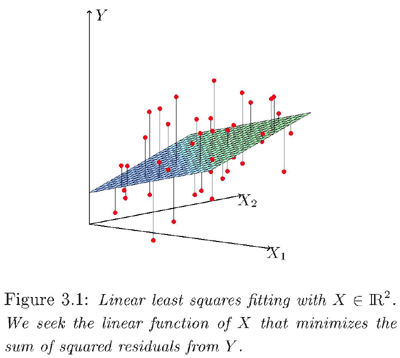
For accurate prediction, hopefully the data will lie close to this hyperplane, but they won't lie exactly in the hyperplane (unless perfect prediction is achieved). In the plot above, the red points are the actual data points. They do not lie on the plane but are close to it.
![]() How should we choose this hyperplane?
How should we choose this hyperplane?
We choose a plane such that the total squared distance from the red points (real data points) to the corresponding predicted points in the plane is minimized. Graphically, if we add up the squares of the lengths of the line segments drawn from the red points to the hyperplane, the optimal hyperplane should yield the minimum sum of squared lengths.
The issue of finding the regression function E(Y | X) is converted to estimating βj , j = 0, 1, ..., p.
Remember in earlier discussions we talked about the trade-off between model complexity and accurate prediction on training data. In this case, we start with a linear model, which is relatively simple. The model complexity issue is taken care of by using a simple linear function. In basic linear regression, there is no explicit action taken to restrict model complexity. [Although variable selection, which we cover in Lesson 4, can be considered a way to control model complexity.]
With the model complexity under check, the next thing we want to do is to have a predictor that fits the training data well.
Let the training data be:
{(x1, y1), (x2, y2), ..., (xN, yN )}, where xi = (xi1, xi2, ..., xip).
Denote β = (β0, β1, ..., βp)T .
Without knowing the true distribution for X and Y, we cannot directly minimize the expected loss.
Instead, the expected loss E(Y − f(X))2 is approximated by the empirical loss RSS(β)/N :
\( \begin {align}RSS(\beta)&=\sum_{i=1}^{N}\left(y_i - f(x_i)\right)^2 \\ &=\sum_{i=1}^{N}\left(y_i - \beta_0 -\sum_{j=1}^{p}x_{ij}\beta_{j}\right)^2 \\ \end {align} \)
This empirical loss is basically the accuracy you computed based on the training data. This is called the residual sum of squares, RSS.
The x's are known numbers from the training data.
Here is the input matrix X of dimension N × (p +1):
\begin{pmatrix}
1 & x_{1,1} &x_{1,2} & ... &x_{1,p} \\
1 & x_{2,1} & x_{2,2} & ... &x_{2,p} \\
... & ... & ... & ... & ... \\
1 & x_{N,1} &x_{N,2} &... & x_{N,p}
\end{pmatrix}
Earlier we mentioned that our training data had N number of points. So, in the example where we were predicting the number of doctors, there were 101 metropolitan areas that were investigated. Therefore, N =101. Dimension p = 3 in this example. The input matrix is augmented with a column of 1's (for the intercept term). So, above you see the first column contains all 1's. Then if you look at every row, every row corresponds to one sample point and the dimensions go from one to p. Hence, the input matrix X is of dimension N × (p +1).
Output vector y:
\[ y= \begin{pmatrix}
y_{1}\\
y_{2}\\
...\\
y_{N}
\end{pmatrix} \]
Again, this is taken from the training data set.
The estimated β is \(\hat{\beta}\) and this is also put in a column vector, (β0, β1, ..., βp).
The fitted values (not the same as the true values) at the training inputs are
\[\hat{y}_{i}=\hat{\beta}_{0}+\sum_{j=1}^{p}x_{ij}\hat{\beta}_{j}\]
and
\[ \hat{y}= \begin{pmatrix}
\hat{y}_{1}\\
\hat{y}_{2}\\
...\\
\hat{y}_{N}
\end{pmatrix} \].
For instance, if you are talking about sample i, the fitted value for sample i would be to take all the values of the x's for sample i, (denoted by xij) and do a linear summation for all of these xij's with weights \(\hat{\beta}_{j}\) and the intercept term \(\hat{\beta}_{0}\).
\(\hat{\beta} =(X^{T}X)^{-1}X^{T}y \)
\(\hat{y} =X\hat{\beta}=X(X^{T}X)^{-1}X^{T}y \)
\(H=X(X^{T}X)^{-1}X^{T} \)
Each column of X is a vector in an N-dimensional space (not the p+1 dimensional feature vector space). Here, we take out columns in matrix X, and this is why they live in N dimensional space. Values for the same variable across all of the samples are put in a vector. I represent this input matrix as the matrix formed by the column vectors:
X = (x0, x1, ..., xp)
Here x0 is the column of 1's for the intercept term. It turns out that the fitted output vector \(\hat{y}\) is a linear combination of the column vectors xj, j = 0, 1, ... , p. Go back and look at the matrix and you will see this.
This means that \(\hat{y}\) lies in the subspace spanned by xj , j = 0, 1, ... , p.
The dimension of the column vectors is N, the number of samples. Usually the number of samples is much bigger than the dimension p. The true y can be any point in this N dimensional space. What we want to find is an approximation constraint in the p+1 dimensional space such that the distance between the true y and the approximation is minimized. It turns out that the residual sum of squares is equal to the square of the Euclidean distance between y and \(\hat{y}\).
\[RSS(\hat{\beta})=\parallel y - \hat{y}\parallel^2 \]
For the optimal solution, \(y-\hat{y}\) has to be perpendicular to the subspace, i.e., \(\hat{y}\) is the projection of y on the subspace spanned by xj , j = 0, 1, ... , p.
Geometrically speaking let's look at a really simple example. Take a look at the diagram below. What we want to find is a \(\hat{y}\) that lies in the hyperplane defined or spanned by x1 and x2. You would draw a perpendicular line from y to the plane to find \(\hat{y}\). This comes from a basic geometric fact. In general, if you want to find some point in a subspace to represent some point in a higher dimensional space, the best you can do is to project that point to your subspace.
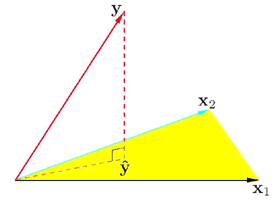
The difference between your approximation and the true vector has to be perpendicular to the subspace.
The geometric interpretation is very helpful for understanding coefficient shrinkage and subset selection (covered in Lessons 4 and 5).
Let's take a look at some results for our earlier example about the number of active physicians in a Standard Metropolitan Statistical Area (SMSA - data available on the Welcome page [1]). If I do the optimization using the equations, I obtain these values below:
\(\hat{Y}_{i}= –143.89+0.341X_{i1}–0.019X_{i2}+0.254X_{i3} \)
\(RSS(\hat{\beta})=52,942,438 \)
Let's take a look at some scatter plots. We plot one variable versus another. For instance in the upper left-hand plot we plot the pairs of x1 and y. These are two-dimensional plots, each variable plotted individually against any other variable.
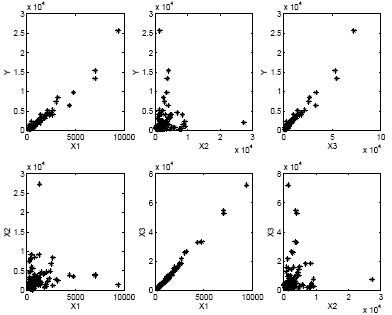
STAT 501 on Linear Regression goes deeper into which scatter plots are more helpful than others. These can be indicative of potential problems that exist in your data. For instance, in the plots above you can see that x3 is almost a perfect linear function of x1. This might indicate that there might be some problems when you do the optimization. What happens is that if x3 is a perfect linear function of x1, then when you solve the linear equation to determine the β's, there is no unique solution. The scatter plots help to discover such potential problems.
In practice, because there is always measurement error, you rarely get a perfect linear relationship. However, you might get something very close. In this case the matrix, XTX, will be close to singular, causing large numerical errors in computation. Therefore, we would like to have predictor variables that are not so strongly correlated.
Here is some theoretical justification for why we do parameter estimation using least squares.
If the linear model is true, i.e., if the conditional expectation of Y given X indeed is a linear function of the Xj's, and Y is the sum of that linear function and an independent Gaussian noise, we have the following properties for least squares estimation.
\[ E(Y|X)=\beta_0+\sum_{j=1}^{p}X_{j}\beta{j} \]
1. The least squares estimation of \(\beta\) is unbiased,
\(E(\hat{\beta}_{j}) =\beta_j, j=0,1, ... , p \)
2. To draw inferences about \(\beta\), further assume: \(Y = E(Y | X) + \epsilon\) where \(\epsilon \sim N(0,\sigma^2)\) and is independent of X.
\(X_{ij}\) are regarded as fixed, \(Y_i\) are random due to ε.
The estimation accuracy of \(\hat{\beta}\), the variance of \(\hat{\beta}\) is given here:
\[Var(\hat{\beta})=(X^{T}X)^{-1}\sigma^2 \]
You should see that the higher \(\sigma^2\) is, the variance of \(\hat{\beta}\) will be higher. This is very natural. Basically if the noise level is high, you're bound to have a large variance in your estimation. But then of course it also depends on \(X^T X\). This is why in experimental design, methods are developed to choose X so that the variance tends to be small.
Note that \(\hat{\beta}\) is a vector and hence its variance is a covariance matrix of size (p + 1) × (p + 1). The covariance matrix not only tells the variance for every individual \(\beta_j\), but also the covariance for any pair of \(\beta_j\) and \(\beta_k\), \(j \ne k\).
This theorem says that the least squares estimator is the best linear unbiased estimator.
Assume that the linear model is true. For any linear combination of the parameters \(\beta_0 , \cdots ,beta_p\) you get a new parameter denoted by \(\theta = a^{T}\beta\). Then \(a^{T}\hat{\beta}\) is just a weighted sum of \(\hat{\beta}_0, ..., \hat{\beta}_p\) and is an unbiased estimator since \(\hat{\beta}\) is unbiased.
We want to estimate θ and the least squares estimate of θ is:
\( \begin {align} \hat{\theta} & = a^T\hat{\beta}\\
& = a^T(X^{T}X)^{-1}Xy \\
& \doteq \tilde{a}^{T}y, \\
\end{align} \)
which is linear in y. The Gauss-Markov theorem states that for any other linear unbiased estimator, \(c^Ty\), the linear estimator obtained from the least squares estimation on \(\theta\) is guaranteed to have a smaller variance than \(c^Ty\):
\[Var(\tilde{a}^{T}y) \le Var(c^{T}y). \]
Keep in mind that you're only comparing with linear unbiased estimators. If the estimator is not linear, or is not unbiased, then it is possible to do better in terms of squared loss.
\(\beta_j\), j = 0, 1, ..., p are special cases of \(a^T\beta\), where \(a^T\) only has one non-zero element that equals 1.
Diabetes data
The diabetes data set is taken from the UCI machine learning database repository at: https://archive.ics.uci.edu/ml/datasets/Pima+Indians+Diabetes [2].
Save the data into your working directory for this course as "diabetes.data." Then load data into R as follows:
# set the working directory
setwd("C:/STAT 897D data mining")
# comma delimited data and no header for each variable
RawData <- read.table("diabetes.data",sep = ",",header=FALSE)
In RawData, the response variable is its last column; and the remaining columns are the predictor variables.
responseY <- RawData[,dim(RawData)[2]]
predictorX <- RawData[,1:(dim(RawData)[2]-1)]
In order to fit linear regression models in R, lm can be used for linear models, which are specified symbolically. A typical model takes the form of response ~ predictors where response is the (numeric) response vector and predictors is a series of predictor variables.
Take the full model and the base model (no predictors used) as examples:
fullModel <- lm(responseY~predictorX[,1]+predictorX[,2]
+predictorX[,3]+predictorX[,4]+predictorX[,5]
+predictorX[,6]+predictorX[,7]+predictorX[,8])
baseModel <- lm(responseY~1)
For the full model, \$coefficients shows the least square estimation for \(\hat{\beta}\) and \$fitted.values are the fitted values for the response variable.
fullModel\$coefficients
fullModel\$fitted.values
Links:
[1] https://onlinecourses.science.psu.edu/stat857/intro
[2] https://archive.ics.uci.edu/ml/datasets/Pima+Indians+Diabetes