

| Key Learning Goals for this Lesson: |
Textbook reading: Consult Course Schedule |
So in linear regression, the more features ($X_j$) the better (since RSS keeps going down)? NO!
Carefully selected features can improve model accuracy. But adding too many can lead to overfitting:
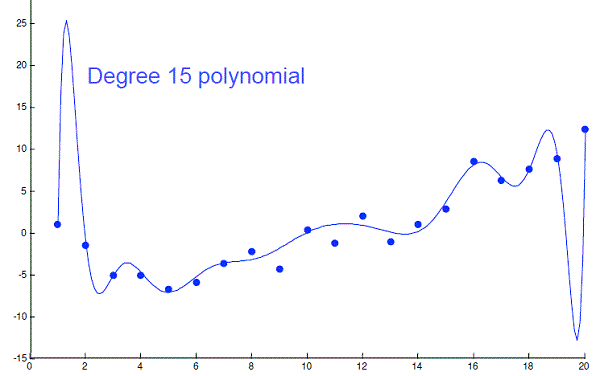
In such cases we would select a subset of predictor variables to perform regression or classification, e.g. to choose k predicting variables from the total of p variables yielding minimum \(RSS(\hat{\beta})\).
When the association of Y and \(X_j\) conditioning on other features is of interest, we are interested in testing \(H_0 : \beta_j= 0\) versus \(H_a : \beta_j \ne 0\).
When prediction is of interest:
The residual sum-of-squares \(RSS(\beta)\) is defined as:
\[RSS(\beta)=\sum_{i=1}^{N}(y_i-\hat{y}_i)^2 = \sum_{i=1}^{N}(y_i-X_i\beta)^2\]
Let \(RSS_1\) correspond to the bigger model with \(p_1 + 1\) parameters, and \(RSS_0\) correspond to the nested smaller model with \(p_0 + 1\) parameters.
The F statistic measures the reduction of RSS per additional parameter in the bigger model:
\[F=\frac{(RSS_0-RSS_1)/(p_1-p_0)}{RSS_1/(N-p_1-1)}\]
Under the normal error assumption, the F statistic will have a \(F_{(p_1-p_0), (N-p_1-1)}\) distribution.
For linear regression models, an individual t-test is equivalent to an F-test for dropping a single coefficient \(\beta_j\) from the model.
Let \(L_1\) be the maximum value of the likelihood of the bigger model.
Let \(L_0\) be the maximum value of the likelihood of the nested smaller model.
The likelihood ratio \(\lambda = L_{0} / L_{1} \) is always between 0 and 1, and the less likely are the restrictive assumptions underlying the smaller model, the smaller will be \(\lambda\).
The likelihood ratio test statistic (deviance), \(-2log(\lambda)\), approximately follows a \(\chi_{p_1-p_0}^{2} \) distribution.
So we can test the fit of the 'null' model \(M_0\) against a more complex model \(M_1\).
Note that the quantiles of the \(F_{(p_1-p_0), (N-p_1-1)}\) distribution approach those of the \(\chi_{p_1-p_0}^{2} \) distribution.
Use of the LRT requires that our models are nested. Akaike (1971/74) proposed a more general measure of "model badness:"
\[AIC=-2 log L(\hat{\beta}) + 2p \]
where p is the number of parameters.
Faced with a collection of putative models, the 'best' (or 'least bad') one can be chosen by seeing which has the lowest AIC.
The scale is statistical, not scientific, but the trade-off is clear; we must improve the log likelihood by one unit for every extra parameter.
AIC is asymptotically equivalent to leave-one-out cross-validation.
AIC tends to overfit models (see Good and Hardin Chapter 12 for how to check this).
Another information criterion which penalizes complex models more severely is:
\[BIC=-2 log L(\hat{\beta})+p\times log(n)\]
also known as the Schwarz' criterion due to Schwarz (1978), where an approximate Bayesian derivation is given.
Lowest BIC is taken to identify the 'best model', as before.
BIC tends to favor simpler models than those chosen by AIC.
An exhaustive search for the subset may not be feasible if p is very large. There are two main alternatives:
There are various methods developed to choose the number of predictors, for instance the F-ratio test. We stop forward or backward stepwise selection when no predictor produces an F-ratio statistic greater than some threshold.
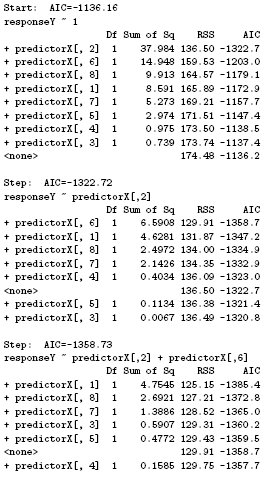
To perform forward stepwise addition and backward stepwise deletion, the R function step is used for subset selection. For forward stepwise selection, baseModel indicates an initial model in the stepwise search and scope defines the range of models examined in the stepwise search. In the example below, the model starts from the base model and expands to the full model.
step(baseModel,scope=list(upper=fullModel,lower=~1),direction="forward")
The result shows the details for the selected predictor variable in each step. In this case Y ~ X2 + X6 + X1 + X7 + X3 + X8 is the best sequence of variables added using forward selection.

Similarly, backward stepwise deletion of variables can be executed by the following code:
step(fullModel, direction="backward")
It is also possible to use the regsubsets function in the leaps library to perform best subset selection and stepwise selection (this is covered in the R Lab for this lesson).
After completing the reading for this lesson, please finish the Quiz and R Lab on ANGEL (check the course schedule for due dates).