Sample R code for Multiple Regression
Qfit1 <- lm(quality ~ ., data=WWTrain50)
summary(Qfit1)
vif(Qfit1)
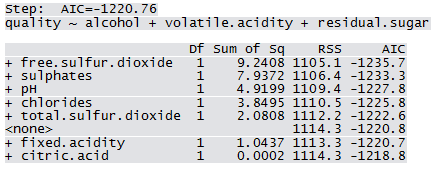
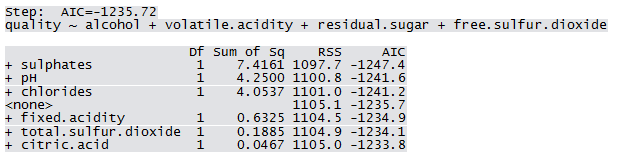
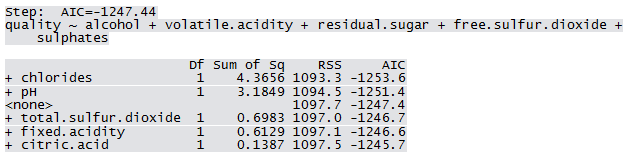
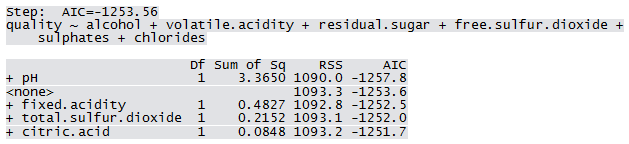
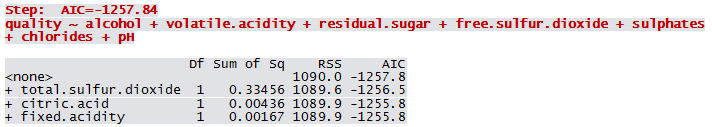
Qfit3 <- step(lm(quality ~ 1, WWTrain50), scope=list(lower=~1, upper = ~fixed.acidity+volatile.acidity+citric.acid+residual.sugar+chlorides+free.sulfur.dioxide+total.sulfur.dioxide+pH+sulphates+alcohol), direction="forward")
Qfit4 <- lm(quality ~ alcohol + volatile.acidity + residual.sugar + free.sulfur.dioxide +
sulphates + chlorides + pH , data=WWTrain50)
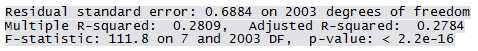
summary(Qfit4)
vif(Qfit4)
Qfit5 <- lm(quality ~ alcohol + volatile.acidity + residual.sugar + rt.sulfur.dioxide +
sulphates + chlorides + pH , data=WWTrain50)
summary(Qfit5)
vif(Qfit5)
par(mfrow=c(1,2), oma = c(3,2,3,2) + 0.1, mar = c(1,1,1,1) + 0.1)
truehist(residuals(Qfit4), h = 0.25, col="slategray3")
qqPlot(residuals(Qfit4), pch=19, col="darkblue", cex=0.6)
mtext("Distribution of Residuals", outer=T, side=1, line = 2)par(mfrow=c(1,1))
pred.val <- round(fitted(Qfit4))
plot(pred.val, residuals(Qfit4))
ts.plot(residuals(Qfit4))
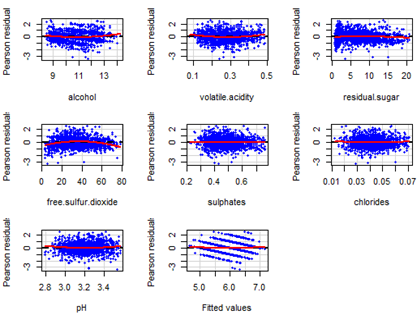
residualPlots(Qfit4, pch=19, col="blue", cex=0.6)
influencePlot(Qfit4, id.method="identify", main="Influence Plot", sub="Circle size is proportial to Cook's Distance" )
boxcox(lm(quality~alcohol), data=WWTrain50, lambda=seq(-0.2, 1.0, len=20), plotit=T)
std.del.res<-studres(Qfit4)
truehist(std.del.res, h = 0.25, col="slategray3")
mtext("Histigram of Studentized Deleted Residuals", side=1, line=2, cex=0.8)d.fit <- dffits(Qfit4)
truehist(std.del.res, h = 0.25, col="slategray3")
truehist(d.fit, h = 0.25, col="slategray3")
mtext("Histigram of Studentized Deleted Residuals", side=1, line=2, cex=0.8)cook.d <- cooks.distance(Qfit4)
ts.plot(cook.d)
par(mfrow=c(1,2), oma = c(3,2,3,2) + 0.1, mar = c(1,1,1,1) + 0.1)
truehist(std.del.res, h = 0.55, col="slategray3")
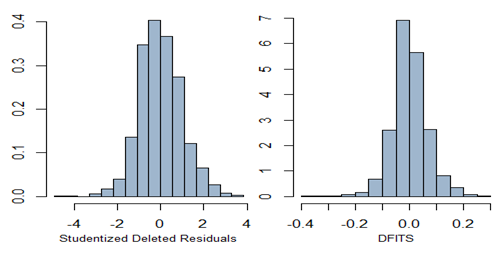
mtext("Studentized Deleted Residuals", side=1, line=2, cex=0.8)truehist(d.fit, h = 0.05, col="slategray3")
mtext("DFITS", side=1, line=2, cex=0.8)par(mfrow=c(1,1), oma = c(3,2,3,2) + 0.1, mar = c(1,1,1,1) + 0.1)
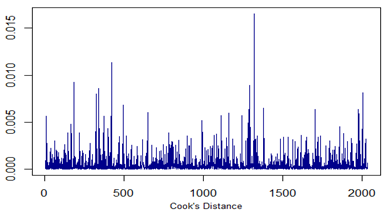
ts.plot(cook.d, col="dark blue")
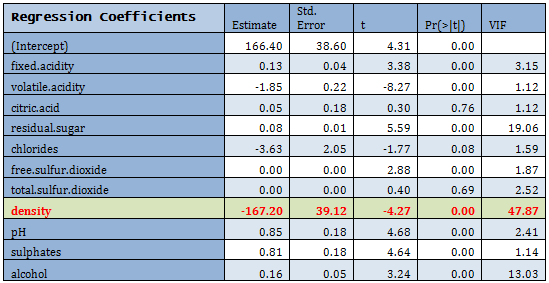
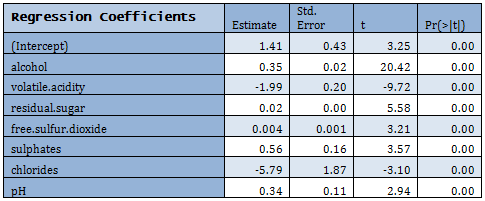
Linear regression is fitted to the Training data.
For extremely high VIF density was removed from the model. There are other predictors with high VIF, but they were not removed at this step.
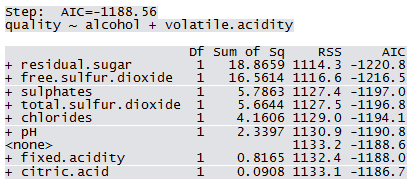
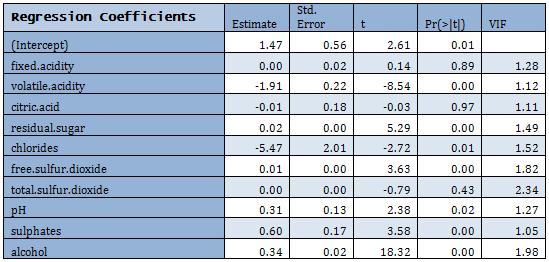
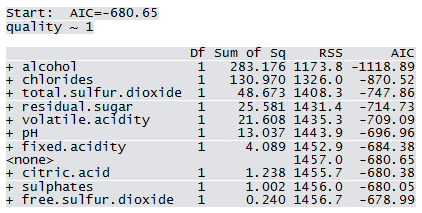
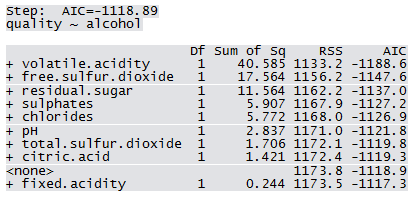
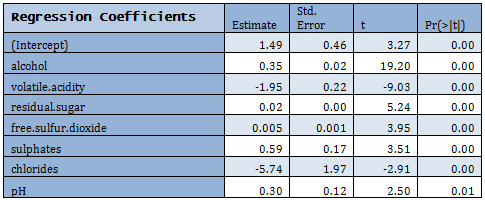
Not all predictors are significant. A forward selection method is employed to build a working model. The sample R output follows:
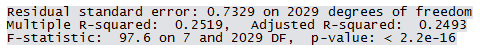
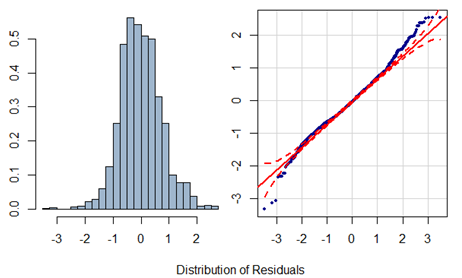
Residuals have an approximately symmetric distribution but there seems to be outliers at both ends. Partial residual plots are given below. Note the pattern in the fitted value plot. Since the response actually takes only integer values but has been assumed to be continuous, such pattern arises.
All three plots are given below. Note that no point is identified as outlier with DFITS value.
Only 26 points are identified as outliers according to the above criteria. A final model is fit after eliminating these points and a slight improvement in the R2 value is noted.
Application of this model on test data gives sum of square of differences between the actual response and predicted response to be 1196.205 whereas sum of square of deviations of actual response is 1554.754. Ratio of these two may be taken as the ratio of Error sum of squares and total sum of squares. Hence a measure similar to that of R2 may be computed as 1 – 1196.205/1554.754 = 0.2306.
Sample R code for Final Model
Qfit6 <- lm(quality ~ poly(alcohol,2) + poly(volatile.acidity,2) + residual.sugar + poly(free.sulfur.dioxide,2) + chlorides + sulphates + poly(pH,2), data=WWTrain50In)
summary(Qfit6)
residualPlots(Qfit6, pch=19, col="blue", cex=0.6)