CD.5: Decision Tree
R codes for
Tree Based Algorithms
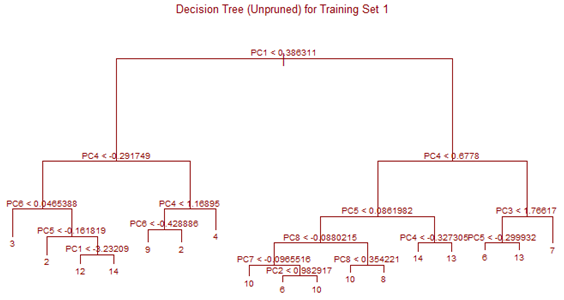
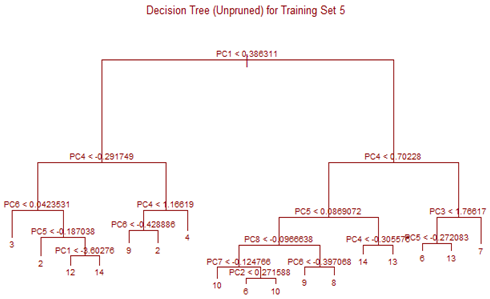
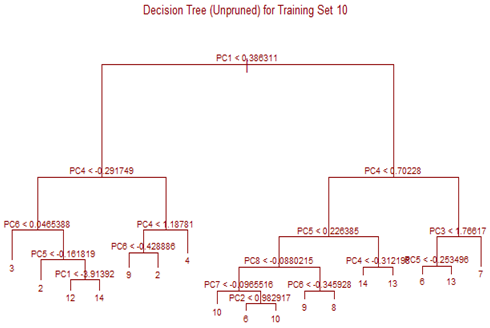
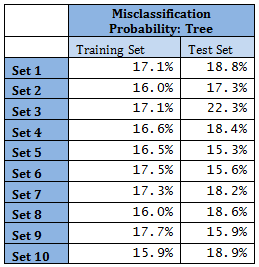
Unsupervised tree algorithm is applied to all Training sets and misclassification probability was calculated for both the Training and Test sets. All the Training Sets give rise to very similar decision trees. Three representative trees are shown below as examples. Following table summarizes the misclassification probabilities for Tree classification Therefore overall mis-classification probability of the 10-fold cross-validation is 17.9%, which is the mean mis-classification probability of the Test sets. Pruning was tried for this decision tree, but it did not improve the result. At the first glance this high error rate compared to k-NN and LDA looks surprising. LDA uses only linear classifier all over the sample space, but Tree procedure recursively partitions the sample space to reduce mis-classification error. It is therefore expected that Tree procedure will always give better results than LDA. However, it is to be noted that LDA takes into account linear combinations of the predictors, whereas Tree always divides the sample space into splits parallel to the axes. If separation is along any other line, Tree wil not be able to capture that. This is exactly what is happening here.